| Gene/ORF
R06F6.1 (cdl-1)
|
Ace View of R06F6.1
|
| R06F6.1 Details |
 |
| Description |
cdl-1 encodes a homolog of human hairpin (stem-loop) binding proteins (HBP/SLBP) that bind to the hairpin (stem-loop) structure in the 3' untranslated region (3' UTR) of histone mRNAs, and thus promote histone pre-mRNA processing and translation of mature histone mRNA; CDL-1 is required for normally high levels of histone gene expression, normal cell division during late larval development, embryonic viability, normal vulval morphogenesis, normally rapid apoptosis, and fertility; CDL-1 binds to the stem-loop structure in the 3' UTR of core-histone mRNA; the cdl-1 promoter is most active in dividing cells during embryogenesis and postembryonic development; both CDL-1 and human HBP contain a minimal RNA-binding domain (RBD) of roughly 73 amino acids that has no similarity with other known RNA-binding motifs. |
| Homology Group (KOG) |
Histone mRNA stem-loop binding protein
|
| GO Terms |
apoptosis
(GO:0006915) ; embryonic development
(GO:0009790) ; mitotic chromosome condensation
(GO:0007076) ; nucleus
(GO:0005634) ; cytoplasm
(GO:0005737) ; mRNA 3' UTR binding
(GO:0003730) |
| Location |
II:10784963..10787580 |
| RNAi Phenotypes |
|
| External Links |
   |
| Associated RNAi Experiments |
View Details
|
| Experiments with Non-Wild Phenotypes |
| Simmer:R06F6.1 |
Emb |
| Cenix:200-b11 |
Emb ; Sister Chromatid Separation abnormal (Cross-eyed) |
| PF:GL1_8E5 |
Delayed P0 spindle rotation ; Ectopic cleavage furrows ; Emb ; EMS extends too far anteriorly ; Excessive blebbing ; Nuclei reform next to cell division remnant ; P0 spindle positioning defect |
| JA:R06F6.1 |
Bmd ; Emb ; Gro ; Rup |
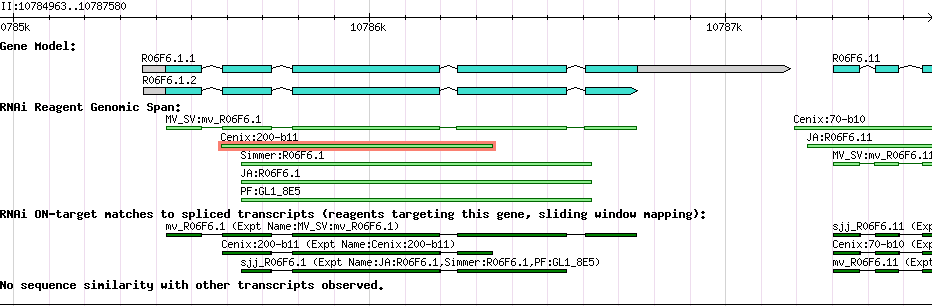
| Gene Plot of R06F6.1 |
Browse using GBrowse
|
|









