| RNAi Experiment: JA:Y18D10A.10 |
Ace View of JA:Y18D10A.10
|
| Genes Inhibited |
 |
| Gene Inhibited |
Y18D10A.10 |
| Description of "Y18D10A.10" |
Associated GO Term : sugar binding |
| KOG Annotation |
C-type lectin
|
 |
| Evidence for Inhibition |
| ePCR |
Status: Unique_ePCR (I:12859481..12861172)
Transcript overlap: 714 nt |

|
| Sliding Window |
Raw score: 630
Relative score: 1 (This gene has the best score for this RNAi reagent)
Specificity index: 0.818 |
| Wormbase (BLAST) |
RNAi primary |
|
| External "Y18D10A.10" Links |
  |
| Gene Inhibited |
T07C4.9 (nex-2) |
| Description of "T07C4.9" |
nex-2 encodes an annexin, a member of a family of calcium-dependent phospholipid binding proteins; by homology, NEX-2 could function in a number of processes, such as membrane fusion, cytoskeletal interactions, and intracellular signaling; however, as loss of nex-2 function via RNA-mediated interference (RNAi) does not result in any abnormalities, the precise role of NEX-2 in C. elegans development and/or behavior is not yet known. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 60
Relative score: 0.095 (Best raw score= for Gene )
Specificity index: 0.078 |
|
| External "T07C4.9" Links |
  |
| Gene Inhibited |
Y51F10.5 |
| Description of "Y51F10.5" |
Associated GO Term : beta-N-acetylhexosaminidase activity |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 80
Relative score: 0.127 (Best raw score= for Gene )
Specificity index: 0.104 |
|
| External "Y51F10.5" Links |
  |
| JA:Y18D10A.10 Phenotypes |
|
 |
| Phenotype |
WT
|
| JA:Y18D10A.10 Experiment Details |
Ace View |
| Laboratory |
JA |
| RNAi Reagent & Type |
sjj_Y18D10A.10 (PCR_reagent) |
| Reference |
Fraser AG et al. (2000) Nature "Functional genomic analysis of C. elegans chromosome I by systematic RNA ...." |
| Date |
2000-11-16 |
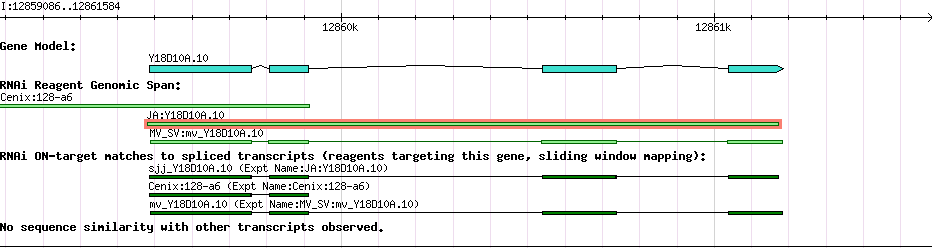
| Gene Plot of Y18D10A.10 |
Browse using GBrowse
|
 |
| Gene Plot of T07C4.9 |
Browse using GBrowse
|
 |
| Gene Plot of Y51F10.5 |
Browse using GBrowse
|
 |














