| RNAi Experiment: Simmer:F17E9.10 |
Ace View of Simmer:F17E9.10
|
| Genes Inhibited |
 |
| Gene Inhibited |
F17E9.10 (his-32) |
| Description of "F17E9.10" |
his-32 encodes an H3 histone; by homology, HIS-32 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-32 is a replication-dependent histone locus that resides in the HIS5 cluster on chromosome IV. |
| Evidence for Inhibition |
| ePCR |
Status: Unique_ePCR (IV:8333982..8334659)
Transcript overlap: 380 nt |

|
| Sliding Window |
Raw score: 338
Relative score: 1 (This gene has the best score for this RNAi reagent)
Specificity index: 0.302 |
| Wormbase (BLAST) |
RNAi primary |
|
| External "F17E9.10" Links |
  |
| Gene Inhibited |
F54E12.1 (his-55) |
| Description of "F54E12.1" |
his-55 encodes an H3 histone; by homology, HIS-55 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-55 is a replication-dependent histone locus that resides in a histone gene-rich region on chromosome IV. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 66
Relative score: 0.195 (Best raw score= for Gene )
Specificity index: 0.059 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F54E12.1" Links |
  |
| Gene Inhibited |
F22B3.2 (his-63) |
| Description of "F22B3.2" |
his-63 encodes an H3 histone; by homology, HIS-63 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-63 is a replication-dependent histone locus. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 75
Relative score: 0.222 (Best raw score= for Gene )
Specificity index: 0.067 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F22B3.2" Links |
  |
| Gene Inhibited |
B0035.10 (his-45) |
| Description of "B0035.10" |
his-45 encodes an H3 histone. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 73
Relative score: 0.216 (Best raw score= for Gene )
Specificity index: 0.065 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "B0035.10" Links |
  |
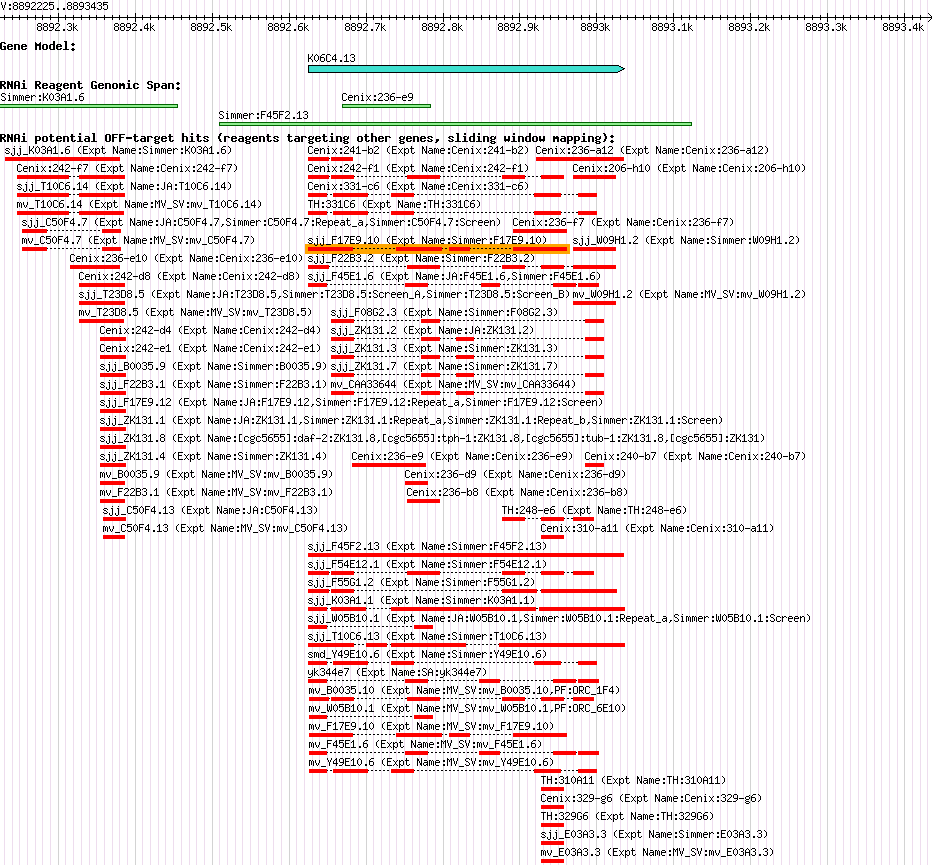
| Gene Inhibited |
K06C4.13 (his-27) |
| Description of "K06C4.13" |
his-27 encodes an H3 histone; by homology, HIS-27 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-27 is a replication-dependent histone locus that resides in the HIS4 cluster on chromosome V. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 84
Specificity index: 0.075 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "K06C4.13" Links |
  |
| Gene Inhibited |
F08G2.3 (his-42) |
| Description of "F08G2.3" |
his-42 encodes an H3 histone. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 62
Relative score: 0.183 (Best raw score= for Gene )
Specificity index: 0.055 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F08G2.3" Links |
  |
| Gene Inhibited |
T10C6.13 (his-2) |
| Description of "T10C6.13" |
his-2 encodes an H3 histone; his-2 is contained within the histone gene cluster HIS1. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 124
Relative score: 0.367 (Best raw score= for Gene )
Specificity index: 0.111 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "T10C6.13" Links |
  |
| Gene Inhibited |
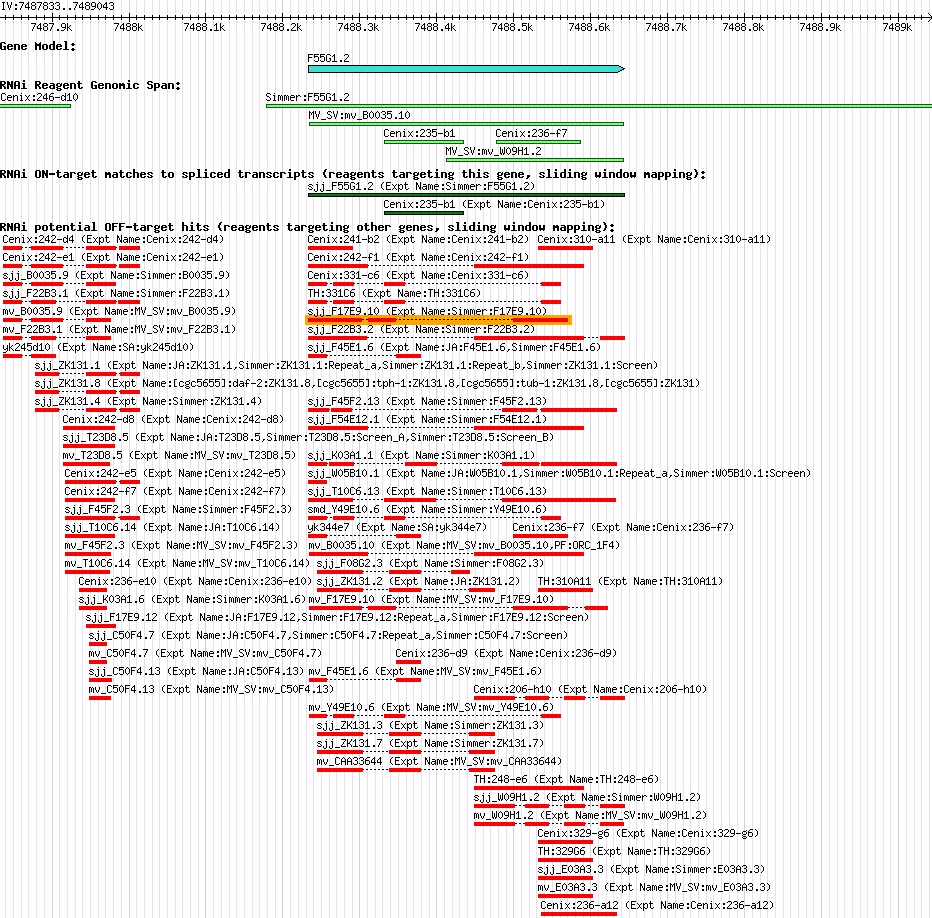
F55G1.2 (his-59) |
| Description of "F55G1.2" |
his-59 encodes an H3 histone; by homology, HIS-59 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-59 is a replication-dependent histone locus that resides in a histone gene-rich region on chromosome IV. |
| KOG Annotation |
Histones H3 and H4
|
 |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 70
Relative score: 0.207 (Best raw score= for Gene )
Specificity index: 0.062 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F55G1.2" Links |
  |
| Gene Inhibited |
F45F2.13 (his-6) |
| Description of "F45F2.13" |
his-6 encodes an H3 histone. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 60
Relative score: 0.178 (Best raw score= for Gene )
Specificity index: 0.054 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F45F2.13" Links |
  |
| Gene Inhibited |
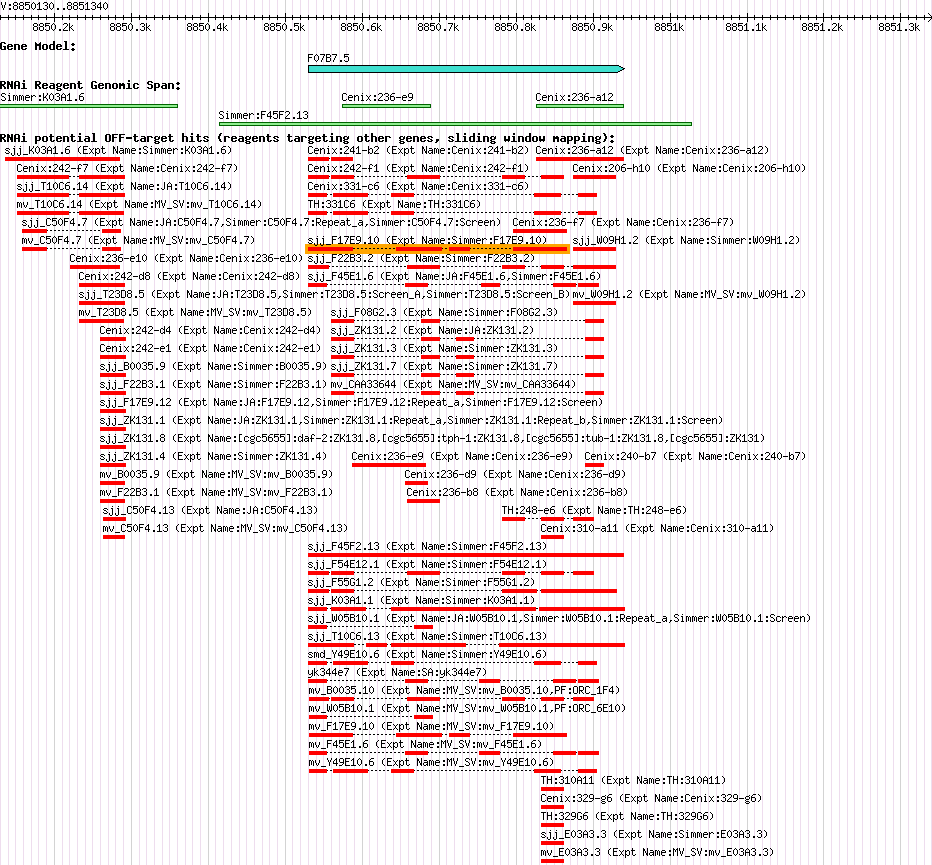
F07B7.5 (his-49) |
| Description of "F07B7.5" |
his-49 encodes an H3 histone; by homology, HIS-49 is predicted to function as a nucleosome component required for packaging of DNA into chromatin; his-49 is a replication-dependent histone locus that resides in a histone gene-rich region on chromosome V. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 84
Specificity index: 0.075 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "F07B7.5" Links |
  |
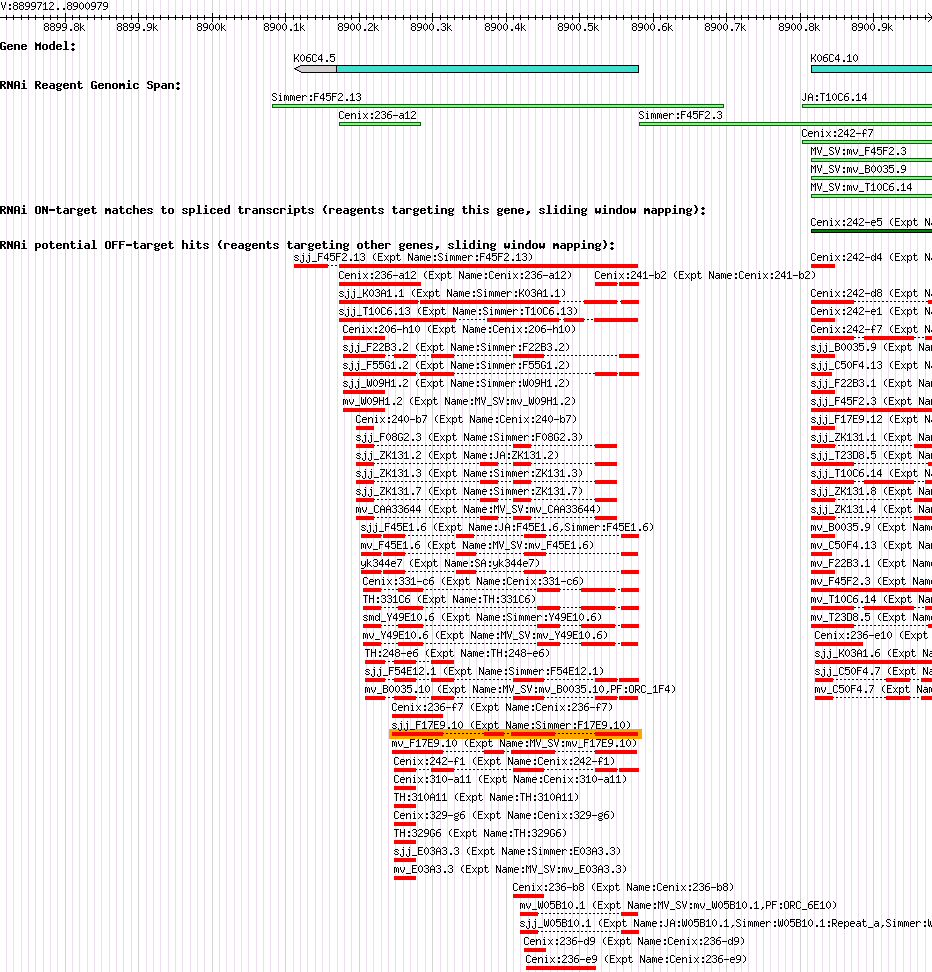
| Gene Inhibited |
K06C4.5 (his-17) |
| Description of "K06C4.5" |
his-17 encodes an H3 histone; his-17 is contained within the histone gene cluster HIS4. |
| Evidence for Inhibition |
| Sliding Window |
Raw score: 84
Relative score: 0.249 (Best raw score= for Gene )
Specificity index: 0.075 |
| Wormbase (BLAST) |
RNAi secondary |
|
| External "K06C4.5" Links |
  |
| Simmer:F17E9.10 Phenotypes |
|
 |
| Embryonic |
Emb Penetrance 50 - 80 % |
| Larval |
Lva
|
| Adult |
Ste
brood size (6 out of 10)
|
| Simmer:F17E9.10 Experiment Details |
Ace View |
| Laboratory |
NL |
| RNAi Reagent & Type |
sjj_F17E9.10 (PCR_reagent) |
| Strain |
NL4256 rrf-3(pk1426) |
| Reference |
Simmer F et al. (2003) PLoS Biology "Genome-wide RNAi of C. elegans using the hypersensitive rrf-3 strain reveals ...." |
| Date |
2003-10-01 |
| Gene Plot of F22B3.2 |
Browse using GBrowse
|
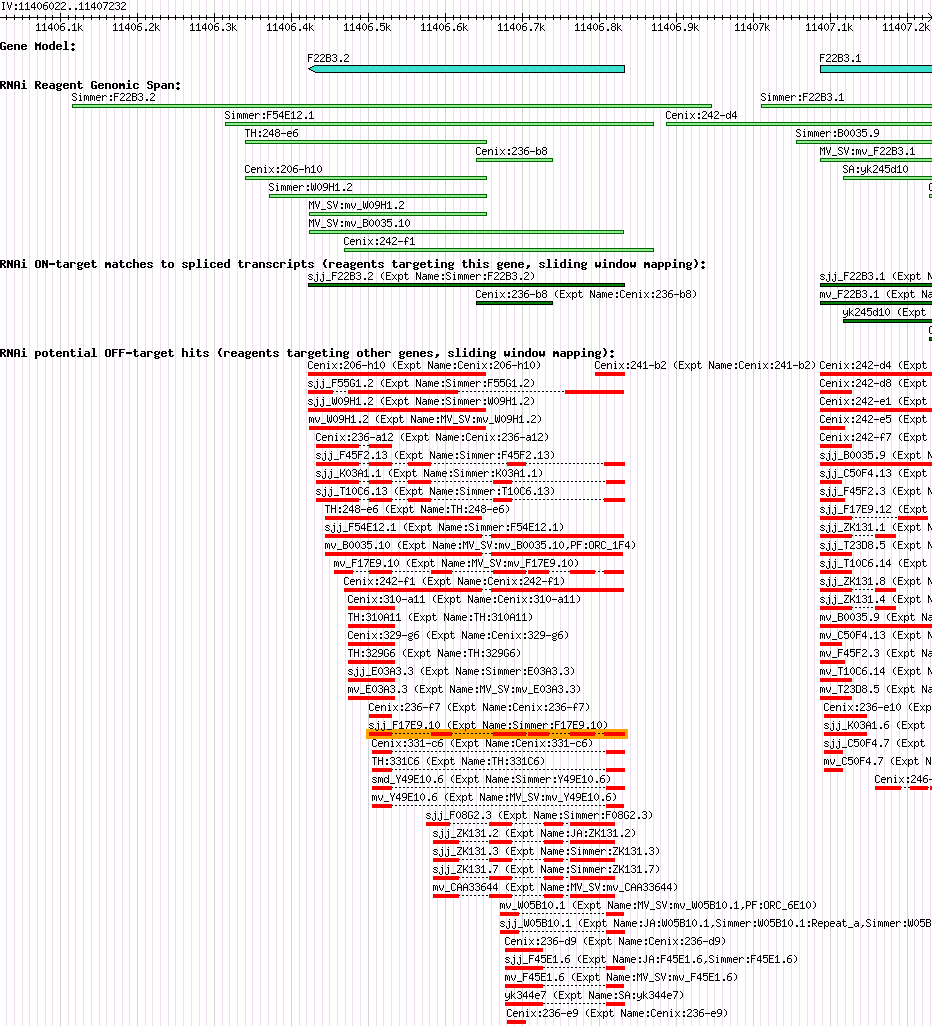 |
| Gene Plot of F54E12.1 |
Browse using GBrowse
|
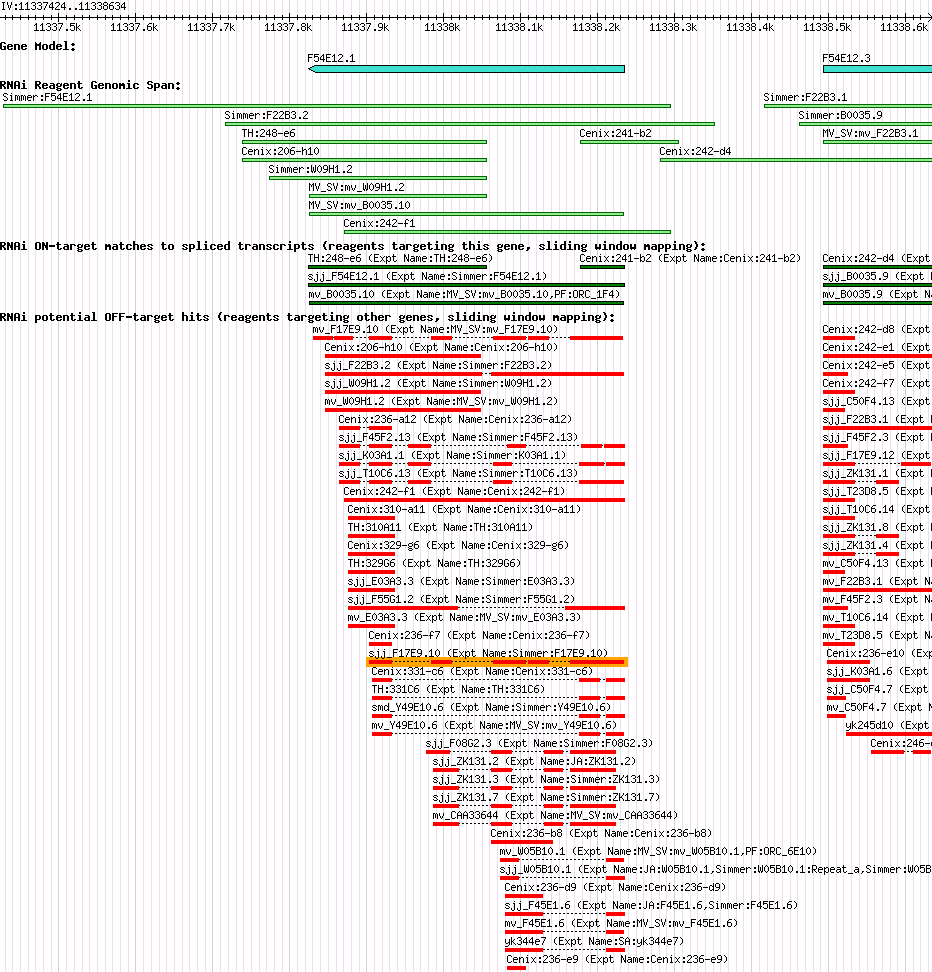 |
| Gene Plot of B0035.10 |
Browse using GBrowse
|
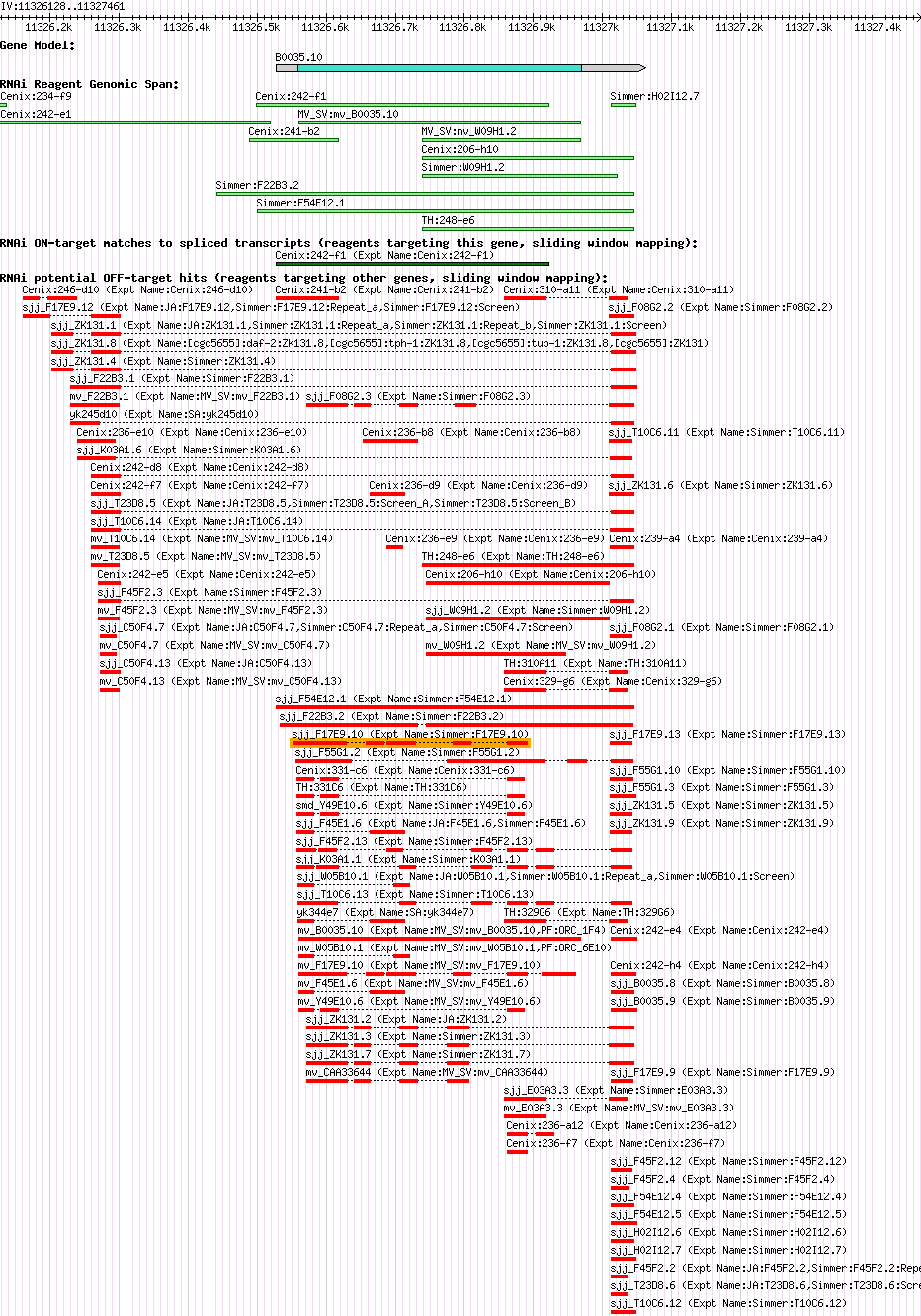 |
| Gene Plot of F17E9.10 |
Browse using GBrowse
|
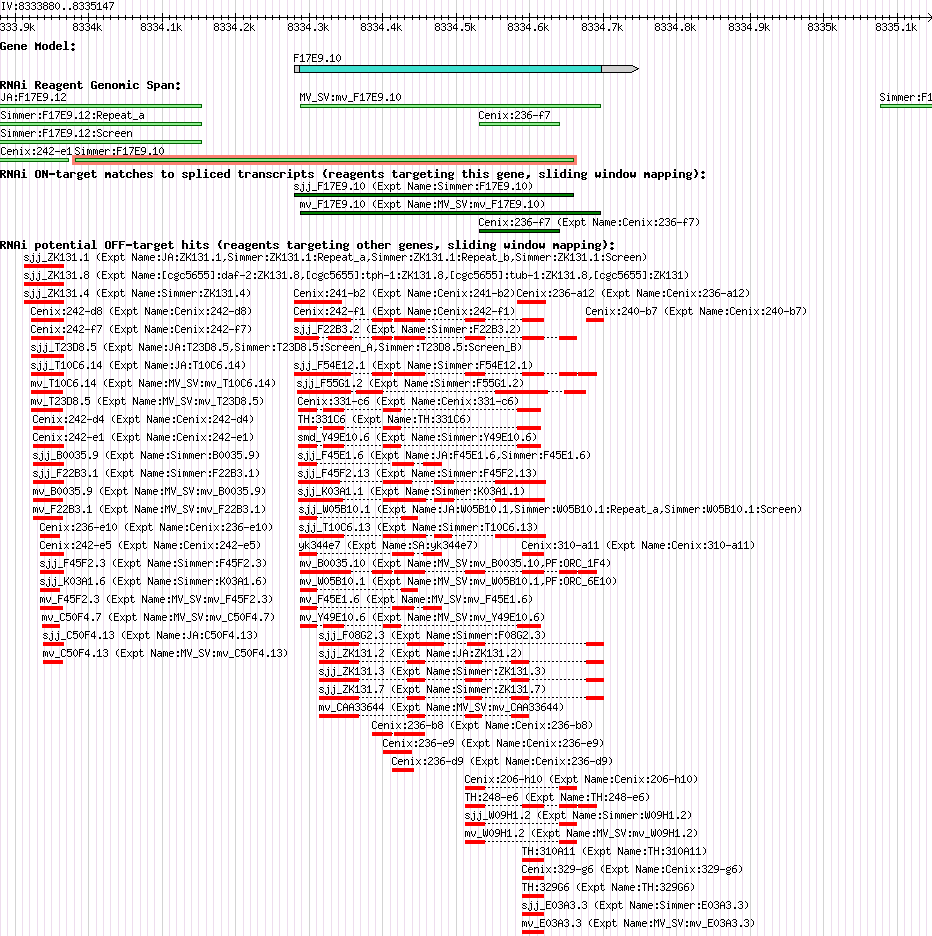 |
| Gene Plot of K06C4.13 |
Browse using GBrowse
|
 |
| Gene Plot of F08G2.3 |
Browse using GBrowse
|
 |
| Gene Plot of T10C6.13 |
Browse using GBrowse
|
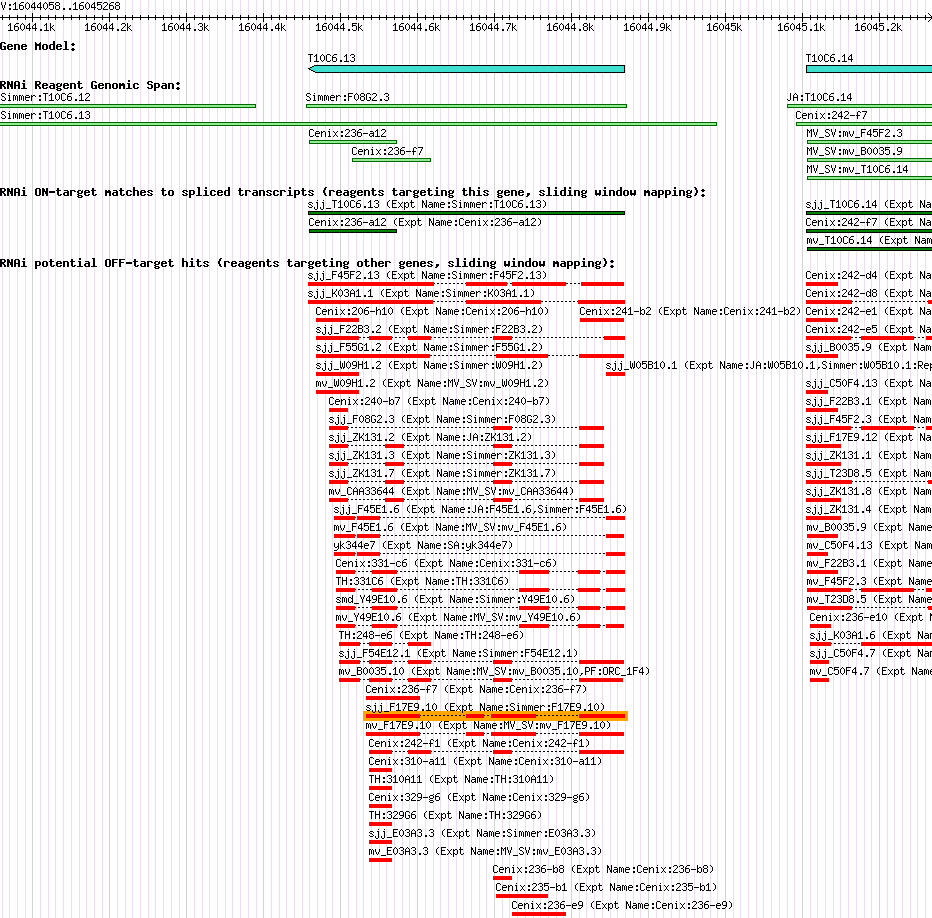 |
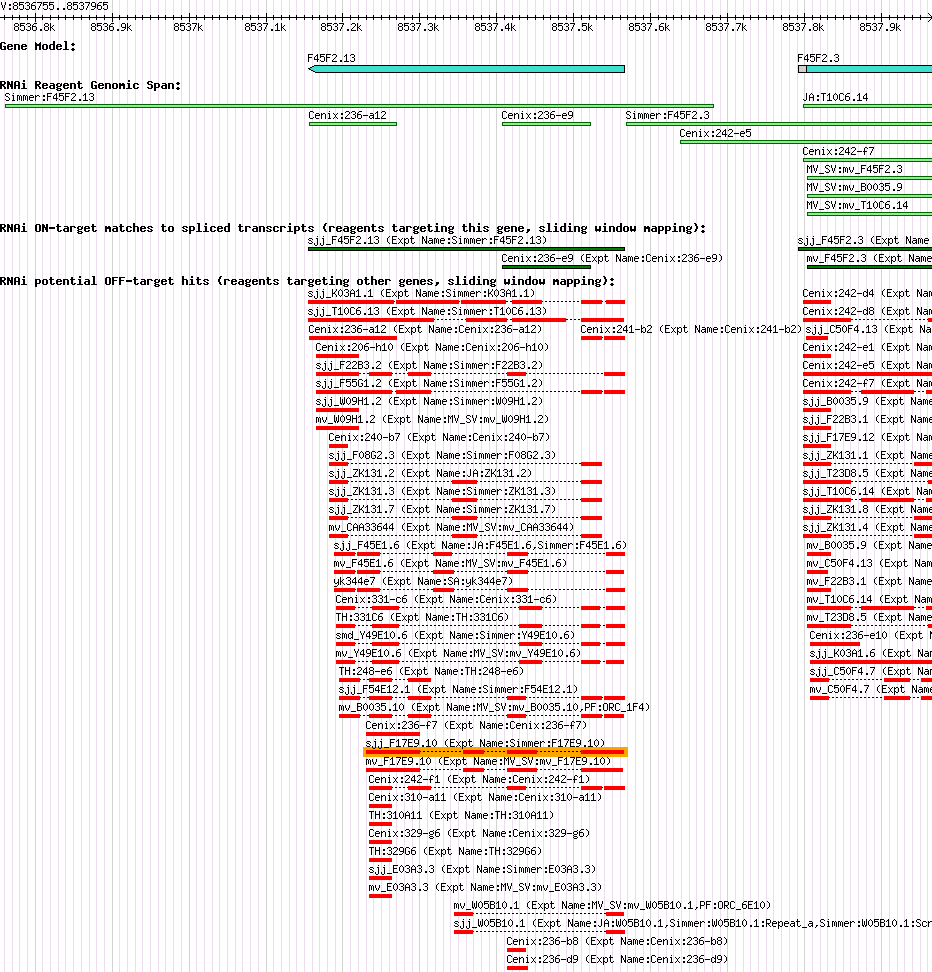
| Gene Plot of F45F2.13 |
Browse using GBrowse
|
 |
| Gene Plot of F55G1.2 |
Browse using GBrowse
|
 |
| Gene Plot of F07B7.5 |
Browse using GBrowse
|
 |
| Gene Plot of K06C4.5 |
Browse using GBrowse
|
 |






































